
Ant colonies are complex systems par excellence. It’s almost as if the colony is the organism, not the ant. Ants follow simple behavioural patterns, depositing pheromones as they go and following trails of scent laid down by others. Because of their collective actions, the colony seems to have a life of its own, sprouting its foraging trails towards food sources much like a slime mold grows its branches along the shortest path to food. The colony appears to have its own reproductive cycle too: queens and males mate during the nuptial flight, and the impregnated queens then land to give birth to new colonies, like fertilized eggs. Ordinary workers play no role in reproduction; they are outside the germline.
But some species of ants behave in ways that are even more complex: they form supercolonies, networks of interconnected nests with hundreds of reproductive queens. In these supercolonies, queens and workers move freely between nests without eliciting aggression; they cooperate across nest boundaries. Ant supercolonies are the largest cooperative units known in nature: for some ants, they can extend for hundreds of kilometres. They are also among the strangest: their existence is difficult to explain from the point of view of gene-centric evolutionary theory. This has to do with altruism: relatedness among nestmates can be low, and workers will end up helping unrelated individuals that carry a different set of genes. It may even be that ant supercolonies represent an evolutionary dead end.
Recently, I had a chance to have some fun with the genetics of ant supercolonies. My colleagues Eva Schultner and Heikki Helanterä who work on ants had collected a number of samples from tens of nests of F. Aquilonia in southern Finland. As Eva and Heikki wanted to understand the genetic structure of F. Aquilonia supercolonies, the sampled ants were genotyped for estimating genetic similarities between the nests (for technical details, scroll down). From a network-science point of view, the nests and their similarities span a weighted spatial network: nests are nodes and pairwise genetic similarities are mapped to link weights. The resulting similarity network looks like this:
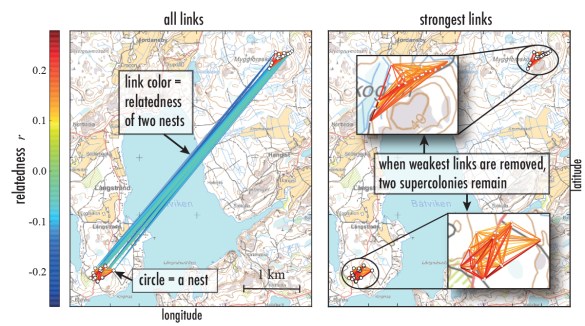
There are two supercolonies, one to the NE and one to the SW – the link weights inside the colonies are higher than between them, much like you would have for two communities in a social network. A closer look inside these two supercolonies (with methods more advanced than bare-bones network thresholding) revealed that there is a faint hint of substructure, of subclusters inside supercolonies. And because queens, workers, and pupae were genotyped separately and sampled at two time points, we could see that the genetic relationships between nests are not the same in terms of queens as they are in terms of workers, and not the same in spring as they are in summer when workers have started migrating.
This means that there may an extra layer of complexity in the genetics of ant supercolonies – fine structure in time and space, and in terms of class.
This work was published in Molecular Ecology last year. If you are interested in toying around with ant genetics, the data are available on Datadryad and my Python scripts can be found here: github.com/jsaramak/ants.
[Technical details: the ants were sequenced at 8 polymorphic microsatellite loci; microsatellites are nonsensical bits of DNA where a random sequence is repeated 5-50 times. They do not do anything and there is no selection pressure, and therefore microsatellite alleles are great for just seeing how close or far two populations are genetically. There are various measures for quantifying this: the simplest would be to see how often the same alleles appear in populations. In social-insect studies, the typical measure is the so-called relatedness (Queller & Goodnight 1989) and we used it in this work.]
Very nice post
LikeLike
Thanks!!
LikeLiked by 1 person
Pingback: Thou Shalt Not Smooth! – Jari Saramäki
Pingback: Do an MSc in Complex Systems – Admission Now Open! – Jari Saramäki